Difference between revisions of "Statistical Manager"
(→Philosophy) |
(→Architecture) |
||
| Line 30: | Line 30: | ||
The subsystem comprises the following components: | The subsystem comprises the following components: | ||
| − | * ''' | + | * '''Ecological Engine Library''': a container for several data mining algorithms and evaluation procedures for the quality assessment of the modeling procedures. Algorithms follow a plug-in implementation and deploy; |
| − | * ''' | + | * '''Computational Infrastructure Occupancy Tree''': an internal process which monitors the occupancy of the resources to choose among when launching an algorithm. |
| − | * ''' | + | * '''Algorithms Thread''': an internal process which puts in connection the Algorithm to execute with a unloaded infrastructure which is able to execute it. Infrastructures are weighted according to the computational speed and the internal logic will choose the fastest available; |
| − | * ''' | + | * '''WS Resource''': an internal gCube process which takes care of all the computations asked by a single user\service. The WS Resource communicates with the other components by means of gCube events; |
| − | * ''' | + | * '''Object Factory''': a broker for WS Resources and a link between the users' computations and the Occupancy Tree process. |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
A diagram of the relationships between these components is reported in the following figure: | A diagram of the relationships between these components is reported in the following figure: | ||
| − | [[Image: | + | [[Image:statistical-manager-architecture.png|frame|center|Statistical Manager Internal Architecture]] |
== Deployment == | == Deployment == | ||
Revision as of 12:17, 9 May 2012
A cross usage service developed in the iMarine Work Package 10, aiming to provide users and services with tools for performing Data mining operations. This document outlines the design rationale, key features, and high-level architecture, as well as the options deployment.
Overview
The goal of this service is to offer a unique access for performing data mining or statistical operations on heterogeneous data. data can reside on client side in the form of csv files or they can be remotely hosted, as SDMX documents or they can be stored in a database.
The Service is able to take such inputs and execute the requested operation by invoking the most suited computational infrastructure, chosing among a set of available possibilities: executions can run on multi-core machines, or on different computational Infrastructure, like the d4Science itself or Windows Azure and other options.
Algorithms are implemented as plug-ins which makes the injection mechanism of new functionalities easy to deploy.
Design
Philosophy
This represents a unique endpoint for those clients or services which want to perform complex operations without going to investigate into the details of the implementation. Currently the set of operations which can be performed has been divided into:
- Generators
- Modelers
- Transducers
- Evaluators
Further details are available at the Ecological Modeling wiki page, where some experiments are shown along with explanations on the algorithms.
Architecture
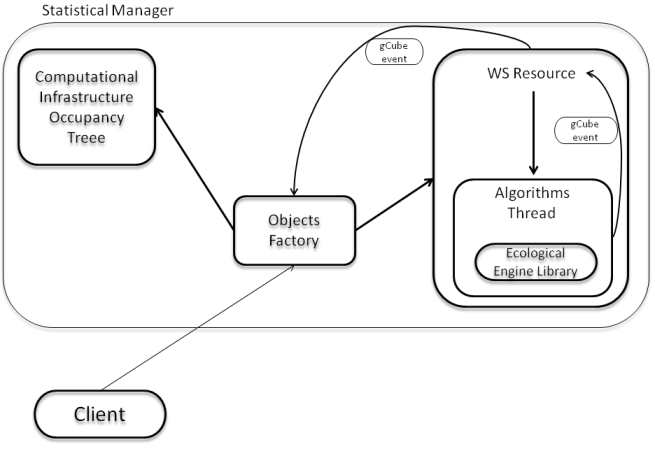
The subsystem comprises the following components:
- Ecological Engine Library: a container for several data mining algorithms and evaluation procedures for the quality assessment of the modeling procedures. Algorithms follow a plug-in implementation and deploy;
- Computational Infrastructure Occupancy Tree: an internal process which monitors the occupancy of the resources to choose among when launching an algorithm.
- Algorithms Thread: an internal process which puts in connection the Algorithm to execute with a unloaded infrastructure which is able to execute it. Infrastructures are weighted according to the computational speed and the internal logic will choose the fastest available;
- WS Resource: an internal gCube process which takes care of all the computations asked by a single user\service. The WS Resource communicates with the other components by means of gCube events;
- Object Factory: a broker for WS Resources and a link between the users' computations and the Occupancy Tree process.
A diagram of the relationships between these components is reported in the following figure:
Deployment
All the components of the subsystem must be deployed together in a single node. This subsystem can be replicated in multiple hosts; this does not guarantee a performance improvement because the scalability of this system depends on the capacity of external repositories contacted by the plugins. There are no temporal constraints on the co-deployment of services and plugins. Every plugin must be deployed on every instance of the service. This subsystem is lightweight, it does not need of excessive memory or disk space.
Small deployment
Use Cases
Well suited Use Cases
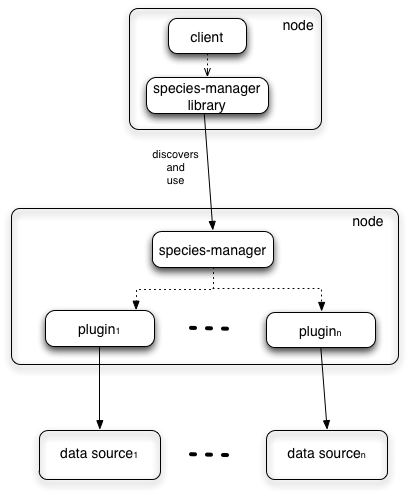
The subsystem is particularly suited to support abstraction over biodiversity data. Every biodiversity repository can be easily integrated in this subsystem developing a plugin.
The development of any plugin of the Species Manager services immediately extends the ability of the systems to discovery new biodiversity data.